The impact of physics on biology and medicine
Feature: September 1999
There is a long tradition of physicists and
physics-based techniques making important contributions to biology and
medicine. Here the director of the National Institutes of Health, one of the
world's foremost biomedical research centers, argues that this tradition must
go on.
The aim of most biomedical research is to
uncover new knowledge that will lead to better health. At the National
Institutes of Health (NIH) in the
In this article, I would like to discuss my
conviction that we can only wage an effective war on disease if the scientific
community harnesses the energies of many disciplines, not just biology and
medicine. These allied disciplines range from mathematics, engineering and
computer science to sociology, anthropology and the behavioral sciences. But
the weight of historical evidence and the prospects for the future place
physics and chemistry most prominent among these disciplines.
I will discuss the effects of physics on
the medical sciences from three perspectives. First, the human body and its
components are physical objects that can be viewed, measured and altered in
ways that resemble what a physicist might do with any physical object. Second,
I will remind you of an enormously important phase in the history of biology in
which physicists transformed the study of living things by helping to discover
the principles of heredity. Third, I will describe
some contemporary problems in the biomedical sciences that I believe present
challenges to physicists, young and old. I will also explain the ways in which
the NIH is attempting to ease the path from a formal training in physics to an
active, investigative role in biomedical sciences.
I am only the latest in a long line of
commentators who have made the really quite obvious point that, for at least
several hundred years, physicists - and especially their principles, methods
and machines - have been illuminating our views of the human body and of every
other living thing.
This notion was brought home to me very
early in life when my father - a general practitioner whose office was directly
connected to our house - showed me how X-rays and fluorography could reveal the
bones and lungs of our pets and his patients, and help make diagnoses of
disease. Röntgen and Edison had been pioneers in this
respect. The significance of using the discoveries of physics to perceive
biological function was further impressed on me at college, when one of my
first independent projects required that I try to explain the repeating peaks
and valleys of my electrocardiogram as a record of voltage changes in the salty
sea of the human body. And at medical school I learned that the doyens of our
biochemistry department had become famous by being the first to tag red blood
cells with easily detected radioisotopes to learn how long such cells survived
in the body.
These few personal memories are just a
sampling of the hundreds of physics-based methods that have been applied to
view living bodies without the disruption of anatomical dissection or to
visualize very small components of living things.
A more
systematic rendering of this topic was offered by the distinguished Stanford
physicist Robert Hofstadter, in a talk to the National Academy of Sciences in
1983 (see table).
It is instructive to note how many of the methods can be classified as
techniques that permit us to visualize the inside of the human body at
successively higher levels of resolution, or allow us to see smaller and
smaller elements of bodily components.
The methods of "macro-imaging"
include conventional X-radiology, computerized tomography scanning, ultrasound,
positron-emission tomography (PET) and magnetic resonance imaging (MRI). The
impact of these procedures on medical practice is unquestioned and continues to
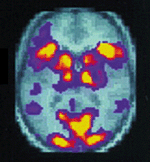
grow as new methods and new applications appear. Two recent examples convey the
exciting potential for both clinical and investigative work - the combined use
of PET and MRI to provide images of the human brain at work (Figure 1), and the
use of MRI to analyze both structural and functional characteristics of the
human heart in diseased states.
"Micro-imaging" began with the
use of optical principles to devise the light microscope, but has progressed too
much higher levels of resolution with electron microscopy, X-ray
crystallography and nuclear magnetic resonance.
Sometimes a collection of methods proves
important, as in the combined use of molecular hybridization, fluorochrome chemistry, wave optics, and computer science in
"spectral karyotyping". This procedure
allows the rapid identification of each of the 23 pairs of normal human
chromosomes and also the origins of recombined chromosomes that often appear in
cancer cells (Figure
2).
Long-awaited success in using a time-honored
technique, X-ray crystallography, to resolve the structure of proteins embedded
in biological membranes has recently transformed the study of cell function and
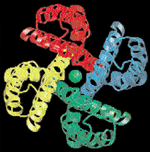
disease. I used an important example of this progress - the analysis by Rod
MacKinnon and co-workers at Rockefeller University in New York (see Doyle et al.
in further reading) of potassium channel proteins to understand how the
channels can be so efficient and yet so selective (Figure 3) - when
justifying further investments in research to Congress this year.
Despite the centrality of such
contributions of physics to modern biology and medicine, I recognize the danger
that my emphasis might be interpreted as limited and perhaps even insulting,
because (some might say) I have portrayed physicists as merely the developers
of tools of measurement that allow biomedical scientists to do the really
important work. There are reasons for my sensitivity to this issue: in a 1967
commentary on the role of physics in biology and medicine, for example, Sergei Feitelberg, a physicist from Mount Sinai Hospital in New
York, noted that while such "spectacular developments created a clear and
unequivocal need for physicists and their help, the role of the physicist was
that of a glorified technician engaged in methodology and instrumentation,
dignified only by the strangeness of his doings and the mysteriousness of his
tools".
I do not accept that interpretation. In
fact, I would argue that we need to show our appreciation of physics-based
technology by investing NIH funds more aggressively in its development. We have
begun to do just that through a new Bioengineering Consortium and a trans-NIH
emphasis on technology development. Still, I would like to address a deeper set
of contributions that physics makes to biology - through the efforts of
physicists who themselves seek to understand the rules of living systems.
Correlations
between physics and medicine
|
Physics |
Medicine |
|
Statics (mechanics) |
Orthopaedics |
|
Dynamics (mechanics) |
Heart motion |
|
Elasticity and strength of
materials |
Orthopaedics |
|
Fluid statics |
Blood pressure |
|
Fluid dynamics |
Blood flow in vascular system |
|
Surface tension |
Capillary action |
|
Sound and acoustics |
Stethoscope, ultrasound,
acoustic microscope |
|
Electricity |
All life processes, ion
transfer at membranes |
|
Magnetism |
Nuclear magnetic resonance
imaging |
|
Light and optics |
Light microscopy, laser
therapy, fiber optics |
|
Heat and thermodynamics |
Heat balance |
|
Kinetic theory and
statistical mechanics |
Brownian motion, osmosis, diffusion
of gases |
|
Atomic physics and
spectroscopy |
"Chemical shift" in
NMR imaging, lasers in medicine |
|
Molecular physics |
Genetics, antibodies, protein
structure, electron microscope |
|
Ultraviolet and infrared
energy |
Skin treatment and imaging |
|
X-rays |
Radiology, CT imaging |
|
Quantum mechanics |
Electron diffraction
microscope |
|
Relativity |
Synchrotron radiation imaging |
|
Crystallography |
Structure of proteins |
|
Solid-state physics and
semiconductors |
Computers in medicine, scintigraphy |
|
Nuclear physics |
Radioisotope labelling, nuclear medicine, radiation therapy |
|
Radioactivity |
Positron emission tomography
(PET) |
|
Elementary particle physics |
Pion
therapy |
|
Accelerators, cyclotrons, etc |
Tumour
therapy, Hodgkin's disease |
|
Astronomy and astrophysics |
Discovery of helium, treatment
of asthma (obsolete) |
This table was presented by Robert
Hofstadter of
Physicists,
heredity and the rise of molecular biology
Exactly 50 years ago, in a speech entitled
"A physicist looks at biology", Max Delbruck, a leading physicist who
had made a conversion to biology some years earlier, attempted to describe the
transition. In the speech, delivered to the 1000th meeting of the Connecticut
Academy of Arts and Sciences, Delbruck said: "A mature physicist,
acquainting himself for the first time with the problems of biology, is puzzled
by the circumstance that there are no 'absolute phenomena'....The animal or
plant or micro-organism he is working with is but a link in an evolutionary
chain of changing forms, none of which has any permanent validity. Even the
molecular species and the chemical reactions which he encounters are the
fashions of today to be replaced by others as evolution goes on. The organism
he is working with is not a particular expression of an ideal organism, but one
thread in the infinite web of all living forms, all interrelated and all
interdependent. The physicist has been reared in a different atmosphere. The
materials and phenomena he works with are the same here and now as they were at
all times and as they are on the most distant star."
Delbruck had been a student of Niels Bohr and then a powerful proselytizer for biology.
With the assistance of Bohr's book Light and Life and, more importantly,
Schrödinger's book What is Life? he attracted many other physicists to biology. The effects
of his missionary zeal were powerful - not just because some very smart people
started to do biology, but because they brought to biological problems a
quantitative, analytic approach - an approach that created the atmosphere in
which principles of molecular biology were discovered by seeking the physical
basis of heredity.
The leading physicist Leo Szilard was among
the converts, and claimed that what physicists brought to biology was "not
any skills acquired in physics, but rather an attitude: the conviction which
few biologists had at that time, that mysteries can be solved" (see Fleming in further
reading).
Delbruck and his friends were gripped by
some fundamental questions: what is the physical form in which hereditary
information is stored? How is it reproduced when a cell divides, or when a
single virus particle invades a cell and makes hundreds or thousands of copies
of itself? How is the information re-assorted during
sexual reproduction? How does the information change when mutations occur?
Answers to many of
these questions, came from the "phage school" that Delbruck founded.
The phage school was a group of former physicists and some biologists who
shared his passion for reducing the problem of heredity to simple rules,
physical entities and conserved energy by studying the replication and genetic
behavior of bacterial viruses (also called bacteriophage
or "phage") in their bacterial hosts. The studies culminated in
findings that form the pillars of modern molecular biology: the identification
of deoxyribonucleic acid (DNA) as genetic material, a description of the
physical organization of DNA through X-ray crystallography, the deduction of
the principles of base pairing and the strategy of replication from the
organization of the double helix, and the deciphering of the genetic code as
triplets chosen from a set of four nucleotides.
Delbruck and his phage school were
important, but there were, in fact, multiple intellectual lineages connected
with physics that helped to create the modern world of molecular biology (see Keller in further
reading). For instance, Warren Weaver was a mathematical physicist turned
science administrator who, in 1932, first used the term "molecular
biology". He chose this phrase because he foresaw "that the moment
would arrive when the distinction between chemistry and physics and even
mathematics on the one hand and biology on the other would be so illusory and
in fact so unfortunate" that he did not want to use the word "biology"
to describe the programs he was supporting at the Rockefeller Foundation.
British scientists with a strong physical
bent, such as Astbury, Bragg and others, used X-ray
diffraction to study the organization of fibres of
many kinds, mainly proteins found in textiles, in an intellectual lineage that
led to Wilkins and Franklin and, of course, DNA. The American geneticists T H
Morgan and H J Muller used physical agents - namely X-rays - to induce
mutations in fruit flies. Muller's affinity for the principles of physics was
especially strong. He was fond of noting the potential similarities of mutation
of genes to transmutation of elements calling the prospect of understanding
these events in physical terms "the two keystones of our rainbow bridges
to power" (see Carlson
in further reading)
Bringing
physics, not just physicists, to biology
To the birth of modern molecular genetics,
physicists contributed their analytic skills but they were not really doing
physics, and many were not even using the computational or imaging tools of
physics as many biologists do. Delbruck and his colleague Salvador Luria
laboriously counted virus infections by hand and eye, just like any other
biologist. But contemporary biology, especially the deciphering of genomes by
nucleotide sequencing, is about to change that. Biology is rapidly becoming a
science that demands more intense mathematical and physical analysis than
biologists have been accustomed to, and such analysis will be required to
understand the workings of cells.
This change was clearly foreshadowed in
Delbruck's 1949 lecture in
But Delbruck also sounded a warning:
"Biology is a very interesting field...[because of] the vastness of its
structure and the extraordinary variety of strange facts...but to the physicist
it is also a depressing subject, because...the analysis seems to have stalled
around in a semi-descriptive manner without noticeably progressing towards a
radical physical explanation...we are not yet at the point where we are
presented with clear paradoxes and this will not happen until the analysis of
the behaviour of living cells has been carried into
far greater detail."
In the past 50 years and especially in the
past 20, molecular and cell biologists have moved much closer to the
"radical physical explanation" of cell behavior that Delbruck sought.
Certainly the chemical elements - especially the genes, the ribonucleic acid
(RNA), and the proteins - and some of their basic functions are coming into
view. What is lacking is a sense of how these functions are integrated to allow
cells to manifest their physiological traits.
I would like to mention three of the
several arenas of biology in which I believe the skills of physicists and their
close cousins can be most productively used.
The first is perhaps the most reductionist.
Methods are now available for examining the physical and chemical properties of
single macromolecules and single complexes of large molecules. These advances
are important because they avoid the need to synchronize a population of molecules
to measure function. Several of these methods and their applications are
reviewed in a special section on "single molecules" in the 12 March
1999 issue of Science. They include laser traps ("optical tweezers")
to study the energetics of molecular motors used for
transport, for contraction and for flagellar motion.
Steven Chu of
Laser traps can also be used to measure the
force of an enzyme complex, such as the one that copies DNA sequences into RNA.
Fluorescence spectroscopy and scanning tunnel microscopy can visualize the
conformation of single large molecules, and methods now in development may soon
be able to determine the order of bases in single, long DNA molecules.
Second, the computational experience of
physical scientists is needed to help interpret complex data sets and the
process of "gene expression". One of the consequences of projects to
sequence the genomes of human beings and many other species is the opportunity
to understand the processes by which the genes of an organism are expressed.
Such information can help us to understand, for example, why some cells develop
into muscle tissue, while others become brain cells. New methods, built on the
availability of a piece of DNA from each gene, allow us to measure the extent
to which genes are read to form RNA (and subsequently protein) in different
tissues and under different environmental conditions.
These micro-methods, called
"expression arrays", are coming into wide use to study bacteria (with
several hundred to a few thousand genes), yeast (with about 6200 genes), worms
(with about 19,100 genes) and vertebrates (which are predicted to contain about
80,000 genes). Some progress has been made through computer-based "cluster
analysis" (see Eisen et al. in further reading) to begin to
interpret the voluminous data that such experiments generate, but biologists
are generally unused to such complex data sets. Recently, I spent an evening at
the Carnegie Institution's observatory at La Serena in
The third area of opportunity for
physicists in biology is the one that most closely approaches Delbruck's goal
of developing a "radical physical explanation" for cell function. In
the past 20 years, mainly through efforts to identify the genes and proteins
that control cell growth and responses to hormones, biomedical investigators
have constructed many so-called signaling pathways that link molecular
interactions at the cell surface to changes in gene expression in the nucleus.
While there is consensus that these linear
pathways are over-simplified, the way forward is far from clear. The pathways
doubtless have many unrecognized components; the information is certainly
flowing between, not just along, the several pathways; and the pathways are
probably regulated in complicated ways through feedback mechanisms and other
means. A few investigators are beginning to grapple with these issues (see Bhalla
and Iyengar, and Weng
et al. in further reading) but there is an obvious need to apply
experiences with potentially analogous complex machines.
Finale:
moving between disciplines
In talking about the effects of one field
on others, I have generally ignored the "boundary problem" - how do
we distinguish among fields? We do this now, in part, by self-identification,
just as we deal with ambiguity about race, ethnicity and religion.
Self-identification in science is commonly linked to the source of one's
graduate degree, and departmental names on diplomas can become limits to
exploration in adjacent fields. But many of us in biology expect that, as
studies of cells and molecules become more obviously in need of several
disciplinary approaches, it will become increasingly difficult to label the
science and to predict the kinds of degrees the people doing it should have.
At the NIH, we have become concerned about
how people should be trained in college and in graduate studies to pursue
biological problems over the next 50 years, and we are discussing the need to
study this issue with the National Research Council. I also agree with Leon
Lederman, who has been leading the movement to establish a more logical order
for teaching the sciences in US high schools: that is physics, chemistry and
then biology. But these activities will come to fruition only after many years,
and it is important to also consider the more immediate need to transport
intellects across artificial disciplinary boundaries.
I sense increasing interest in attempting
to open borders that have been traditionally hard to cross. In the
There are many anecdotal accounts of
successful interdisciplinary training programs. Within our intramural research
program at the NIH, physicists and physics trainees from the US and abroad do
graduate thesis work, take courses in biological topics, and engage in
post-doctoral training that promotes interactions with biologists and
clinicians. Much of this activity occurs under the direction of some of our
most prestigious scientists - such as Ad Bax, Bob Balaban, Bill Eaton and Adrian Parsegian
- and includes work on small-molecule and protein NMR, brain and cardiac MRI,
and other topics, leading to good job prospects for trainees.
On the occasion of the 100th anniversary of
the American Physical Society, I thank physicists for their many contributions
to biology and medicine - for providing the tools that allow us to see and
probe living things, and for training great minds that have uncovered some of
the most fundamental principles of biology. I now encourage physicists to work
collaboratively with biologists as we strive to achieve Delbruck's "radical
physical explanation" for biological systems.
About
the author
Harold Varmus is director of the National
Institutes of Health,
Further reading
U. S. Bhalla and R Iyengar 1999, Emergent
properties of networks of biological signaling pathways Science 283
381-387
E. A. Carlson 1971, An unacknowledged founding of molecular biology: H. J.
Muller's contributions to gene theory, 1910-1936 J. Hist. Biol. 4
160-161
M. Delbruck 1949, A
physicist looks at biology Trans. Conn. Acad. Arts and Sci. 38
173-190
D. A. Doyle et
al. 1998, The structure of the potassium channel: molecular basis of K+
conduction and selectivity Science 280 69-77
B. Eisen et al. 1998, Cluster analysis
and display of genome-wide expression patterns Proc. Natl
Acad. Sci. USA 95 14 863-14 868
S. Feitelberg
1967, Disciplines: physics, biology, and medicine J. Mt Sinai Hosp. NY 34
378-381
D. Fleming 1968, Emigre physicists and the biological revolution Perspect. Amer. Hist. 2 161
R. Hofstadter 1984, Cross strands linking
physics and medicine National Conference on Biological Imaging (National
Academy Press,
E. F. Keller 1990,
Physics and the emergence of molecular biology: a history of cognitive and
political synergy J. Hist. Biol. 23 389-409
G. Weng et al. 1999, Complexity in
biological signaling systems Science 284 92